The World Health Organization (WHO) has designated antibiotic resistance as one of the ten greatest global health threats. Effectively recording, integrating, and utilizing antimicrobial resistance data has become an urgent priority in clinical medicine and scientific research! Dr. Hung-Yi Chiou, Director of the Institute of Population Health Sciences at the National Health Research Institutes; Professor Hsiao-Hui Tsou; and Assistant Chieh Cheng collaborated with infectious disease expert Dr. Shu-Chen Kuo; FHIR format experts Professor Yi-Hsin Yang; Dr. Hsiu-An Lee, and Associate Professor Li-Hui Lee (Figure 1) to develop a Taiwan-specific antibiotic resistance data format template. This template helps systematize and standardize data previously scattered across various stages of the testing process. It not only enhances the accuracy of data recording but also facilitates data exchange between hospitals, further expanding its application value for clinical decision-making and research analysis!
Figure 1. Contributors
If a bacterial infection occurs, how can we determine which antibiotic is effective? In hospitals, a comprehensive antibiotic test involves several crucial steps. Taking the assessment of bacterial resistance in sputum as an example, the process begins with a microscopic examination. This initial step involves observing the specimen (sputum) for the presence of cells (such as epithelial cells), white blood cells, or bacteria. While bacterial characteristics may be visible at this stage, the specific species cannot yet be confirmed. Therefore, the bacteria in the sample are simultaneously cultured and identified. Only after confirming the bacterial species can further testing be conducted to assess its response to various antibiotics—this is the antibiotic susceptibility testing. Based on these results, the doctor selects the most effective and appropriate antibiotic for the patient’s treatment. (Figure 2)
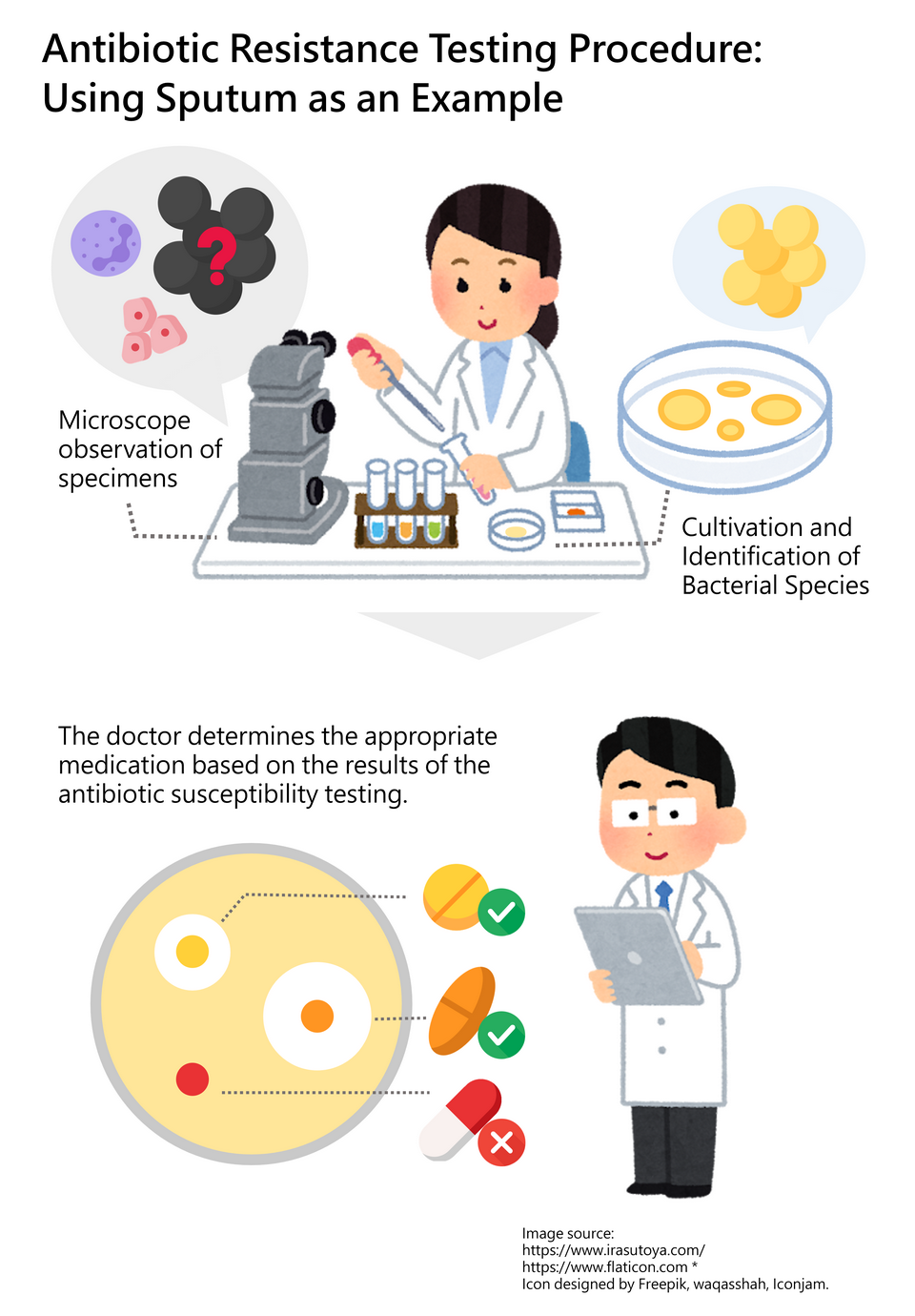
Figure 2. Antibiotic Resistance Testing Process
Clearly documenting each interconnected testing step not only ensures medical accuracy but also enables rapid understanding by other hospitals during referrals. Furthermore, accumulating this data into a long-term database to observe overall trends in antibiotic resistance can provide immense benefits to both patients and society! Therefore, the team adopted the internationally recognized FHIR (Fast Healthcare Interoperability Resources) format as the standard and followed the design specifications outlined in the CDM IG (Common Data Model Implementation Guide) to structure the data.
The FHIR format examples for antibiotic resistance testing are now available on this page. These examples include simulated antibiotic resistance test reports, architectural diagrams, JSON files, and SNOMED CT (Systematized Nomenclature of Medicine — Clinical Terms) code tables for common pathogens. They aim to establish a solid foundation for clinical healthcare and research applications. This example provides simulated antibiotic resistance test reports, architectural diagrams, JSON files, and SNOMED CT (Systematized Nomenclature of Medicine — Clinical Terms) code tables for common pathogens. It aims to establish a solid foundation for clinical healthcare and research applications.
For related materials, please refer to the “Taiwan Infectious Disease Thematic Database: FHIR Zone” managed by the Institute of Population Health Sciences, National Health Research Institutes. https://tidtd.nhri.edu.tw/fhir專區/
